This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Introduction
|
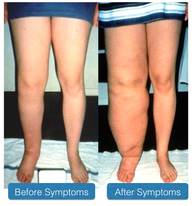
Emberger Syndrome (ES) is a rare autosomal dominant disorder characterized by the co-occurrence of primary lymphedema with a predisposition to acute myeloid leukemia (AML) [1]. Primary lymphedema is the buildup of lymphatic fluid due to missing or impaired lymphatic vessels [2]. Acute myeloid leukemia is a cancer of the blood and bone marrow in which there is an accumulation of abnormal white blood cells, called myeloblasts, that restricts the production of normal blood cells. Additional symptoms that may vary with each case include: warts, sensorineural deafness and minor anomalies such as neck webbing or slender fingers [3].
|
Figure 1: A patient with lymphedema.
|
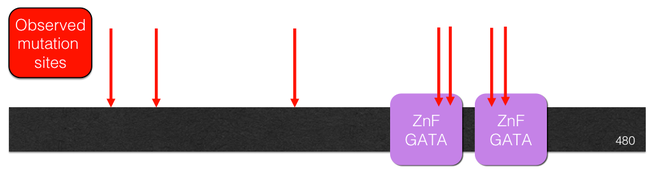
ES is caused by various mutations in the GATA2 gene that result in GATA2 haploinsufficiency, which is when there is only one functional copy of the gene. GATA2 localizes in the nucleus and makes GATA2 protein, which is a transcription factor. GATA2 plays an essential role in regulating the transcription of genes involved in hematopoiesis, which is the formation of blood cellular components, and the regulation of body fluid levels. GATA2 is relatively well-conserved across model organisms, due to it being vital for normal function of the circulatory system. It contains two GATA zinc-finger domains, which bind a specific sequence to activate or repress gene transcription.
Figure 2: The locations of the various mutations in GATA2 found in patients with Emberger Syndrome.
Although research has identified the mechanism of GATA2 haploinsufficiency, how GATA2 regulates the function of genes involved in hematopoiesis remains unclear. The long-term goal of our research is to understand the pathology underlying Emberger Syndrome and GATA2 deficiency. In order to do that, we must first understand the complete role of GATA2 in the development and function of hematopoietic stem cells. Here, we will test the hypothesis that mutations in GATA2 regulate genes that affect the function of hematopoietic stem cells. To test our hypothesis, we will pursue the following aims:
Aim 1: Identify GATA2 interacting proteins
|
First, I wanted to look at the different proteins that GATA2 interacts with, specifically interacting proteins involved in hematopoiesis. I used the STRING database to identify protein interactions, and the gene ontology enrichment option to determine the function of each interacting protein. Since GATA2 is a transcription factor I expected it to have many interacting proteins, specifically proteins involved in processes related to the symptoms observed in ES (lymphedema and AML).
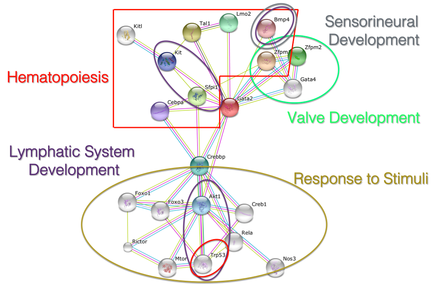
I found that GATA2 interacts with proteins involved in numerous processes, which makes sense given the various phenotypes observed in ES patients. As previously mentioned, some ES patients suffer from deafness, which is likely associated with a disruption in the interaction between GATA2 and Bmp4, which is involved in sensorineural development. |
Figure 3: GATA2 protein interaction network found using STRING database.
|
Most of the interacting proteins were either of regulatory function or developmental function. The group of proteins identified as being involved in the regulation of hematopoiesis are going to be the main focus of my next two aims, as I attempt to look further into the onset of AML. Additionally, there are many interacting proteins involved in the development of vital parts of the circulatory system, such as valves and the lymphatic system. A large collection of proteins are involved in response to stimuli, which are the proteins I will use as my control group going forward.
Aim 2: Identify GATA2 interacting proteins that are differentially expressed in mutant mice
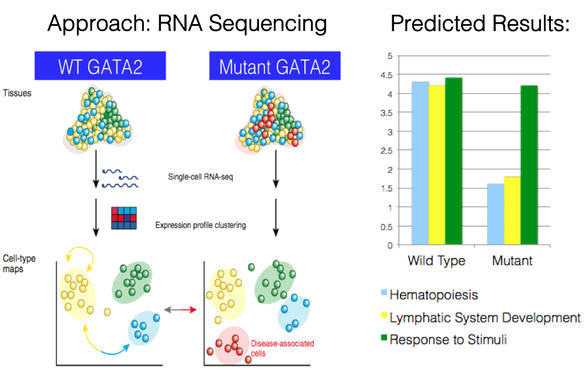
For my second aim I want to determine which of the GATA2 interacting proteins are differentially expressed in mutant mice. I plan to do this by analyzing wild type and mutant mouse bone marrow samples through the use of RNA sequencing. Through RNA sequencing I can look at the expression levels of the genes that produce the interacting proteins. I chose to analyze bone marrow samples because it is involved in hematopoiesis and it also is a key component of the lymphatic system, producing lymphocytes (the fluid that builds up in lymphedema). Since most of the hematopoietic interacting proteins are involved in regulating hematopoiesis, I would expect that they decrease in expression when GATA2 protein is not functioning at a normal level. Similarly, I would expect that the expression levels of genes involved in the lymphatic system development to decrease in expression given the missing/impaired lymphatic vessels that result in lymphedema being observed. Additionally, I would use the group of genes found to respond to stimuli as a control, as I would not expect their expression to be significantly altered.
Figure 4: Experimental approach and expected results for Aim 2.
Aim 3: Determine if differentially expressed GATA2 interacting proteins cause ES phenotypes
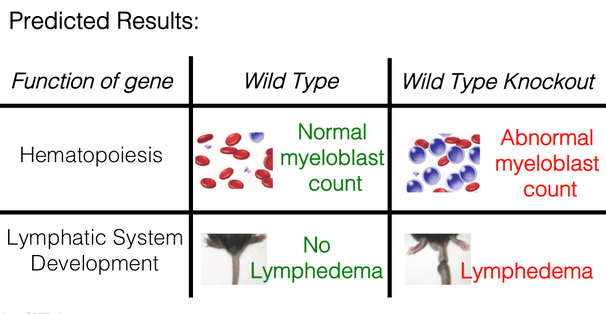
Once I find which genes are differentially expressed, I want to confirm if they cause ES phenotypes. The ES phenotype I am most concerned with is a high myeloblast count, which is observed in AML. I will also look at lymphedema, but this can sometimes be difficult to detect in model organisms. To do this, I plan on taking wild type mice and knocking out the genes found to decrease in expression in mutant mice via the CRISPR/Cas9 technique, while leaving GATA2 fully functional. After knocking out the genes involved in hematopoiesis that were found to be under-expressed in aim 2, I would expect an abnormally high myeloblast count. This would confirm that this set of genes is responsible for the onset of AML in ES patients. Similarly, I would expect the knock out of genes involved in lymphatic system development to result in lymphedema, however these results are likely to be less consistent given the difficulties in observing lymphedema in model organisms.
Figure 5: Predicted results from the CRISPR/Cas9 knock out of differentially expressed genes in wild type mice.
Future Directions
Being able to identify the specific genes responsible for the phenotypes observed in Emberger Syndrome will provide a base that allows for more in-depth studies on the mechanisms behind each phenotype. In the future there is also hope for finding compounds that are capable of binding to various interacting proteins that are differentially expressed due to abnormally functioning GATA2. Finding these compounds could restore function of interacting proteins even without normal GATA2 function. In the future I expect our knowledge to grow on the function of these interacting proteins, as well as GATA2, so that someday we can hopefully find a cure for ES.
References:
1) Mansour, Sahar et al. (2010). Emberger Syndrome – Primary Lymphedema With Myelodysplasia: Report of Seven New Cases.American Journal of Medical Genetics, Part A: 2287-2296.
2) National Lymphedema Network: http://www.lymphnet.org/le-faqs/what-is-lymphedema
3) Ostergaard, Pia et al. (2011). Mutations in GATA2 cause primary lymphedema associated with a predisposition to acute myeloid leukemia (Emberger Syndrome). Nature Genetics, 43(10): 929-930.
Images:
https://asisay.files.wordpress.com/2010/05/lymphedemabeforeafter2.jpg
http://www.fairview.org/healthlibrary/Article/40351
http://jaha.ahajournals.org/content/2/5/e000438/F1.large.jpg
1) Mansour, Sahar et al. (2010). Emberger Syndrome – Primary Lymphedema With Myelodysplasia: Report of Seven New Cases.American Journal of Medical Genetics, Part A: 2287-2296.
2) National Lymphedema Network: http://www.lymphnet.org/le-faqs/what-is-lymphedema
3) Ostergaard, Pia et al. (2011). Mutations in GATA2 cause primary lymphedema associated with a predisposition to acute myeloid leukemia (Emberger Syndrome). Nature Genetics, 43(10): 929-930.
Images:
https://asisay.files.wordpress.com/2010/05/lymphedemabeforeafter2.jpg
http://www.fairview.org/healthlibrary/Article/40351
http://jaha.ahajournals.org/content/2/5/e000438/F1.large.jpg
| Tongas_Final_Presentation_5-5-15.pdf | |
| File Size: | 4704 kb |
| File Type: | |