This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
GATA2 Protein
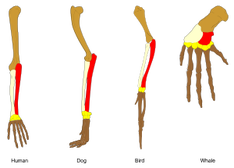
What is Homology?Homology is when different species share similar structures or gene/protein sequences due to common ancestry (1). The image to the right is a simple example that is frequently used to explain homology. As you can see, humans, dogs, birds and whales all share a similar limb structure inherited from a common ancestor. The differences are simply due to evolutionary changes.
|
Figure 1: A comparison of limb structure across four different species that depicts homology.
|
GATA2 gene homology was first determined through the use of the Homologene. This database, however, does not contain information on all species. A good way to determine additional homologs is through the use of BLAST. This tool aligns sequences from a library of genes across species to determine genes that share a significant portion of their sequences. Using BLAST one can determine which species a specific gene is most conserved in. Below are some of the species found to have a gene homologous with human GATA2.
|
Homo sapiens (Humans)
Endothelial transcription factor GATA-2 Accession Number: P23769 |
|
|
Pan troglodytes (Chimpanzee)
GATA binding protein 2 Accession Number: H2RAN3 100% Identical Mus musculus (Mouse)
GATA binding protein 2 Accession Number: O09100 97.9% Identical Cavia porcellus (Guinea Pig)
GATA binding protein 2 Accession Number: H0VFF7 68.5% Identical Loxodonta africana (African elephant)
Uncharacterized Protein Accession Number: G3U3A4 83.4% Identical Gorilla gorilla (Gorilla)
Uncharacterized Protein Accession Number: G3QTJ7 99.2% Identical Macaca mulatta (Rhesus macaque)
Uncharacterized Protein Accession Number: F7FID3 99.8% Identical Pongo abelii (Orangutan)
GATA binding protein 2 Accession Number: H2P9F0 84.8% Identical Danio rerio (Zebrafish) GATA-binding protein 2a Accession Number: Q7T3G1 67.9% Identical Takifugu rubripes (Japanese pufferfish)
Uncharacterized Protein Accession Number: H2TMC1 69.71% Identical Caenorhabditis elegans
Transcription factor elt-1 Accession Number: P28515 71% Identical |
|
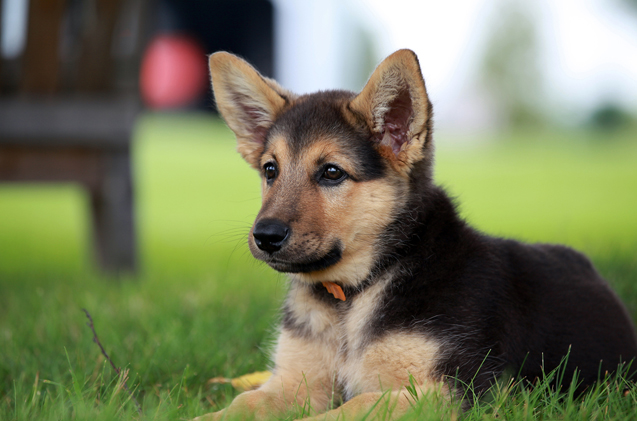
Canis lupus familiaris (Dog)
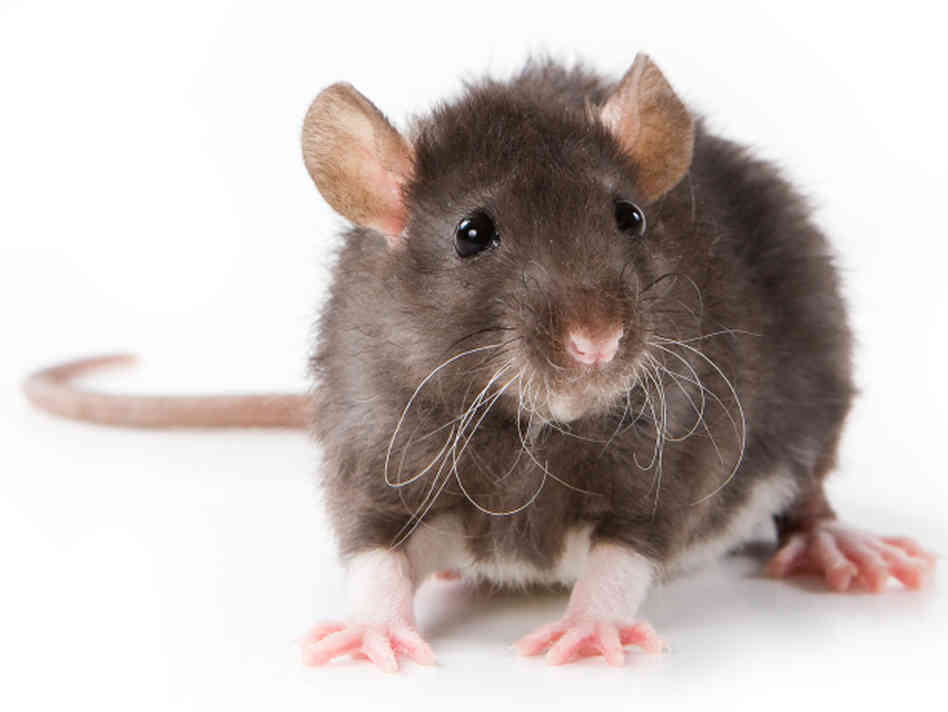
GATA binding protein 2 Accession Number: E2RQ60 98.3% Identical Rattus norvegicus (Rat)
GATA binding protein 2 Accession Number: Q924Y4 97.5% Identical Gallus gallus (Chicken)
GATA binding protein 2 Accession Number: P23824 86.25% Identical Nomascus leucogenys (Gibbon)
Uncharacterized Protein Accession Number: G1RVX7 100% Identical Equus caballus (Horse)
Uncharacterized Protein Accession Number: F6XH29 99% Identical Monodelphis domestica (Gray short-tailed opossum)
Uncharacterized Protein Accession Number: F6ZA02 91.25% Identical Oryctolagus cuniculus (Rabbit)
Uncharacterized Protein Accession Number: G1SUL8 81.2% Identical Felis catus (Cat)
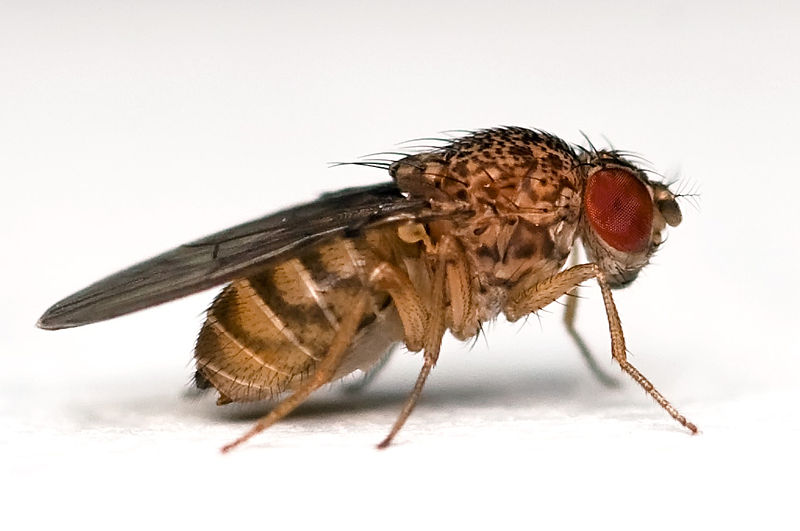
Uncharacterized Protein Accession Number: M3W5E4 98.5% Identical Drosophila melanogaster (Fruit fly)
GATA-binding factor C Accession Number: P91623 54% Identical |
Discussion
The GATA2 protein is, for the most part, well conserved amongst mammals. GATA2 is less conserved in the two invertebrates (C. elegans & D. melanogaster). This could be due to the fact that Drosophila melanogaster is the only species on the list with an open circulatory system and Caenorhabditis elegans is the only species on the list without a circulatory system.
What is Phylogeny?
Phylogenetics is a method commonly used to determine relationships amongst species, genes or proteins (1). These relationships are displayed through phylogenetic trees. There are various components of a phylogenetic tree that each present unique information about the relationships between species, genes or proteins. The number of branches between two species indicates how related they are, with fewer branches suggesting a close relationship and many branches suggesting a distant relationship.
Phylogeny of the GATA2 Protein
The first step in creating a phylogenetic tree is to determine the organisms of interest and to obtain each of their sequences. If you are interested in looking at a specific protein, then you only need the sequence of that protein in FASTA format. After obtaining the different sequences, you must then align them by using one of many programs. For my phylogenetic analysis of GATA2, I used Clustal Omega. After viewing these results using a plug in called Jalview, there are numerous methods that can be applied to form a phylogenetic tree. The methods that I used are described below.
BLOSUM Matrix
This method uses an analysis matrix that looks at how many of the aligned amino acids match and the likelihood of these matches being due to chance in a random sequence (2).
Percent Identity
The second method to compare sequences was using percent identity. As its name suggests, this method simply looks at the percentage of two sequences that are identical. It does this by looking at the number of matching amino acids at each specific location, comparing it to the number of possible amino acids at each location (3).
Neighbor Joining
This method uses the similarity scores obtained from Blosum Matrix or Percent Identity to form a tree that minimizes branch length between two species at a node the more closely related they are (4). The longer a branch, the more change that has occurred
Average Distance
The average distance method also uses the similarity scores obtained from Blosum Matrix or Percent Identity to form a tree, however all branch lengths of closest related species are identical (5). Identical branch lengths suggests equal divergence from a common ancestor.
BLOSUM Matrix
This method uses an analysis matrix that looks at how many of the aligned amino acids match and the likelihood of these matches being due to chance in a random sequence (2).
Percent Identity
The second method to compare sequences was using percent identity. As its name suggests, this method simply looks at the percentage of two sequences that are identical. It does this by looking at the number of matching amino acids at each specific location, comparing it to the number of possible amino acids at each location (3).
Neighbor Joining
This method uses the similarity scores obtained from Blosum Matrix or Percent Identity to form a tree that minimizes branch length between two species at a node the more closely related they are (4). The longer a branch, the more change that has occurred
Average Distance
The average distance method also uses the similarity scores obtained from Blosum Matrix or Percent Identity to form a tree, however all branch lengths of closest related species are identical (5). Identical branch lengths suggests equal divergence from a common ancestor.
|
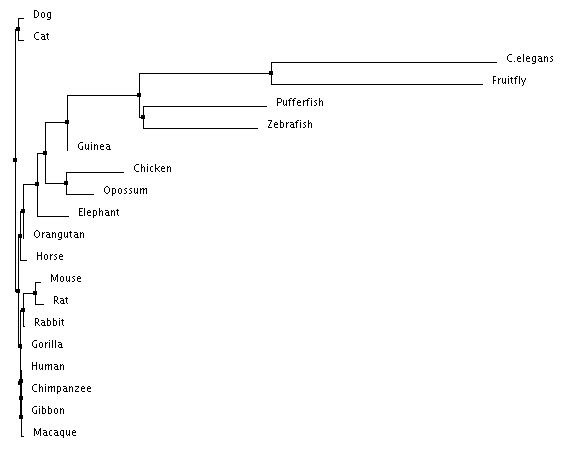
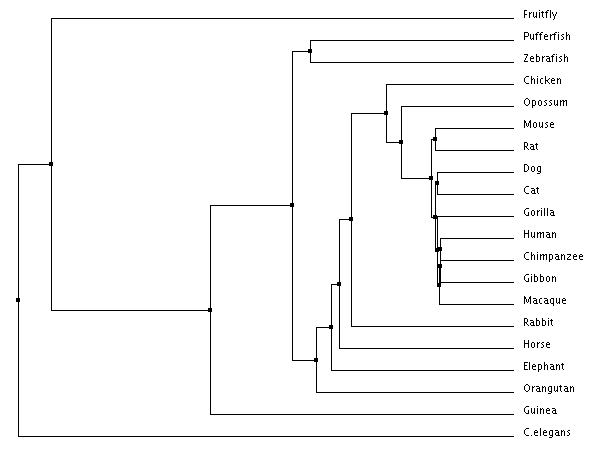
Figure 1: Average distance tree using percent identity
|
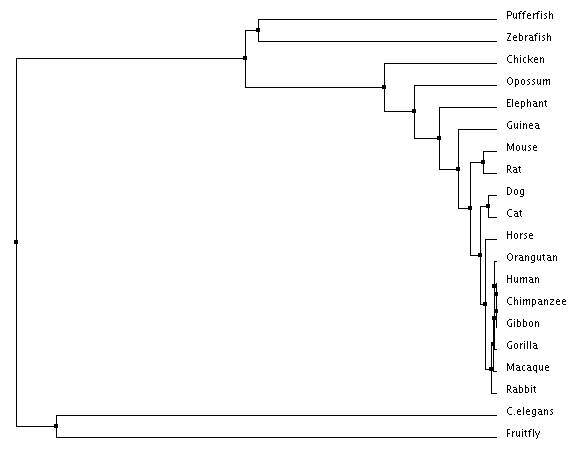
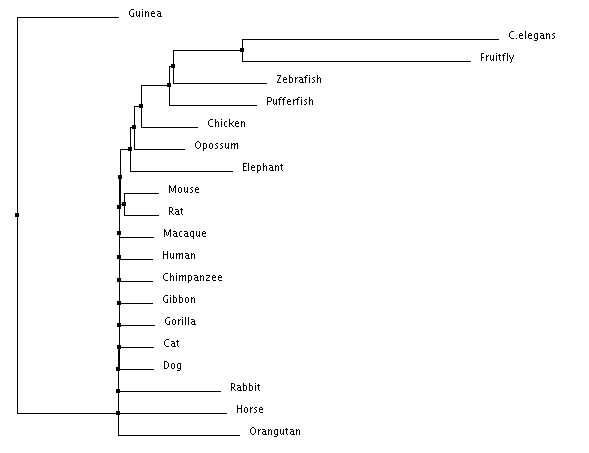
Figure 2: Neighbor joining tree using percent identity
|
Discussion
All trees showed similar relationships between species, with each combination of methods having a few differences. In both average distance trees (percent identity & BLOSUM), C. elegans and Drosophila melanogaster were more distantly related all other species. This makes sense when you look at the role of GATA2 and then consider that C. elegans is the only species on the list without a circulatory system and D. melanogaster is the only species with an open circulatory system. For both of the Neighbor joining trees the most distantly related species are all mammals, which is interesting considering what was previously stated about C. elegans and D. melanogaster.
References:
1) Delsuc, Frederic et al. (2005). Phylogenomics and the reconstruction of the tree of life. Nature Reviews Genetics, 6: 361-375.
2) Substitution Matrices available in Jalview: http://www.jalview.org/help/html/calculations/scorematrices.html
3) Calculations of trees from alignments: http://www.jalview.org/help/html/calculations/tree.html
4) Saitou N and Nei M (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology and Evolution, 4(4): 402-425.
5) Genetics 564 - Lab 2, Homology and Phylogeny: http://genetics564.weebly.com/homology--phylogeny.html
Images (in order of appearance):
http://en.wikipedia.org/wiki/Homology_%28biology%29
http://blog.heartland.org/wp-content/uploads/2014/02/barack-obama-smiling.jpg
http://www.pets4homes.co.uk/images/articles/962/large/6-large-domestic-cat-breeds-with-wild-relatives-51eff964e9d7b.jpg
http://assets.worldwildlife.org/photos/5057/images/featured_story/orangutan_with_baby.jpg?1378992357
http://www.petguide.com/wp-content/uploads/2013/05/cute-dog-names-11.jpg
http://voices.nationalgeographic.com//files/2013/07/69369-cb1373915625-3.jpg
https://www.modernpest.com/wp-content/uploads/2013/10/Photo2-Mouse-2.jpg
http://arlingtonva.s3.amazonaws.com/wp-content/uploads/sites/25/2013/12/rat.jpg
http://www.lovemyguineapig.com/images/looking-guinea-pig2.jpg
http://web.stanford.edu/~jay/gorillas/Jock,_the_Gorilla.jpg
http://t1.gstatic.com/images?q=tbn:ANd9GcRVOX0sy-bZ0F_E7O6zgMEoG0neTKSi1D-V_S4Hga8FKlRbdb0W0zAl_RYi
http://t0.gstatic.com/images?q=tbn:ANd9GcT_gSdBfjJfzR95TSr_DjBTbHa5TaT5AjcfPg5ojo-wqY4Pc7Ih82_pCg
http://t0.gstatic.com/images?q=tbn:ANd9GcQ_-9PGXwxtqvD5ai7u9v7HRdV7NlhTh4PshCEh6Sx3KOlxKhNjH6KHhM8
http://t1.gstatic.com/images?q=tbn:ANd9GcTgJeLDabpBQa8cmSmgFX97rTXGhtTl4rmW5R-_LOurGG1zGNJjlDmM_9UV
http://t3.gstatic.com/images?q=tbn:ANd9GcS7l1QQSAG_5HoED8eReFiYYXSdnfF9pEh4N_CQLzYbcy_mt-pBNyWpxCz2
http://t1.gstatic.com/images?q=tbn:ANd9GcS70J1xdQ6Bc0Yx8LCDd-2eyLsxBtKROGUjBm1vfSyHlazX6vZfLjmwF4oV
http://funmozar.com/rabbit/
http://www.ferris.edu/htmls/colleges/artsands/biological-sciences/faculty-staff/hoerter/zebrafish-model.htm
http://tx.english-ch.com/teacher/nicole/home/the-delicious-but-poisonous-food/
http://lsernafsi2013.weebly.com/methods-and-materials.html
http://en.wikipedia.org/wiki/File:Drosophila_repleta_lateral.jpg