This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Protein Interaction Networks
What are protein interaction networks?
Proteins are complex molecules that are essential for human development and survival. Known, intentional interactions between proteins are displayed in the form of networks. These protein interaction networks revolve around a protein(s) of choice, but are composed of many proteins and can become quite complicated. Using the online database STRING, which is a collection of known and predicted protein interactions, I was able to look at the proteins GATA2 is known to interact with. STRING allows you to form your network by connecting interacting proteins based on either confidence, evidence or action. I was able to use a gene ontology enrichment option to group interacting proteins function.
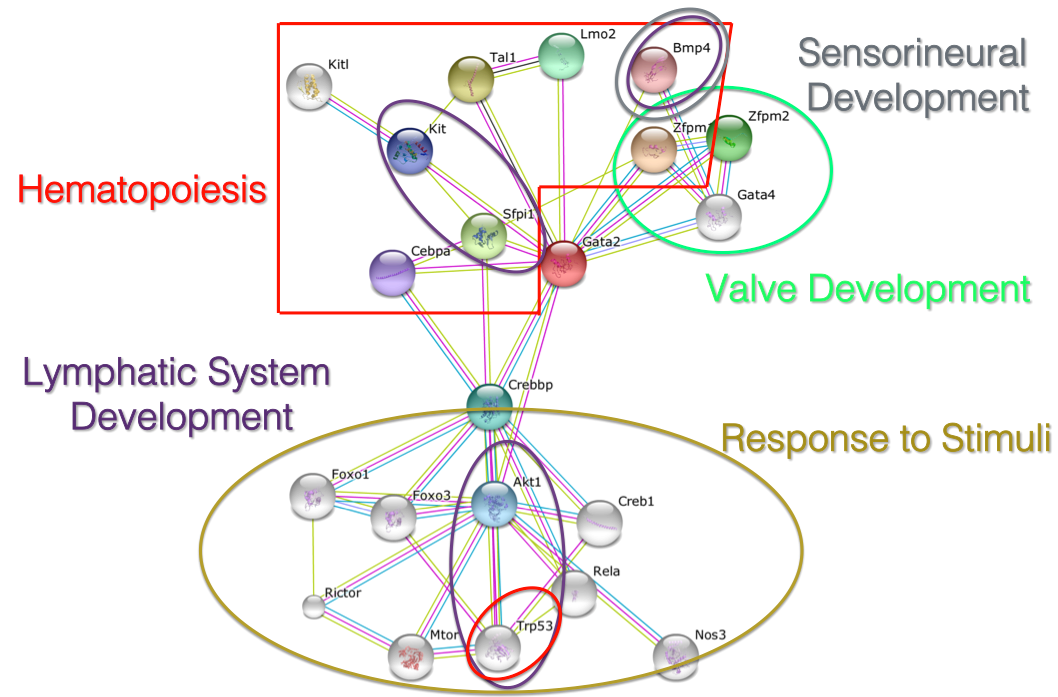
GATA2 interaction network
|
Processes that interacting proteins are involved in:
|
Discussion
Since there is a wide range of symptoms in Emberger Syndrome, I expected GATA2 to interact with a variety of proteins. Interestingly, multiple proteins were found to be involved in a regulatory step of hematopoiesis, including: Kitl, Tal1, Lmo2 and Zpfm1. It is likely that interactions between some of these proteins and GATA2 are disrupted in patients with ES, leading to a buildup of myeloblasts and thus the onset of AML. Proteins found to be involved in the development of the lymphatic system include: Akt1, Trp53, Sfpi1, Kit and BMP4. I expect that changes in the interactions of GATA2 with these proteins results in the damaged lymphatic vessels, ultimately leading to the lymphedema observed in ES patients. Improper interaction between GATA2 and BMP4, which is also involved in sensorineural development, could lead to the deafness that has been observed in some patients. Identifying the function of interacting proteins will allow for more specific studies on individual interactions to determine the role that they play in ES phenotypes.
References:
1) STRING Database: http://string-db.org/